CV for Plants and Animals
Computer vision for plants and animals is an emerging research area that tackles the unique challenges of nature, including complex shapes, diverse appearances, flexible deformations, and highly variable observation conditions. Our work spans foundational problems in visual understanding of plants and animals, from segmenting or modeling plant indivisuals to understanding multimodal features of animal species. These efforts not only open up new directions in computer vision, but also support impactful applications in agriculture, ecology, and biology.
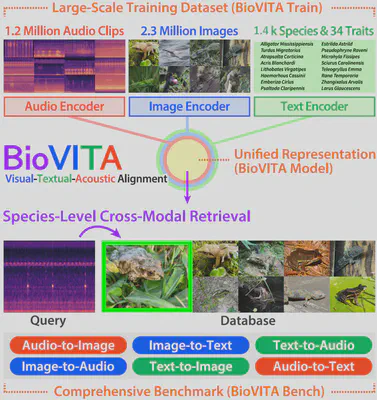
BioVITA: Large-scale Dataset, Model, and Benchmark for Multi-modal Animal Recognition
Understanding animal species from multimodal data poses an emerging challenge at the intersection of computer vision and ecology. While recent biological models, such as BioCLIP, have demonstrated strong alignment between images and textual taxonomic information for species identification, the integration of the audio modality remains an open problem. We propose BioVITA, a novel visual-textual-acoustic alignment framework for biological applications. BioVITA involves (i) a training dataset, (ii) a representation model, and (iii) a retrieval benchmark. First, we construct a large-scale training dataset comprising 1.3 million audio clips and 2.3 million images, covering 14,133 species annotated with 34 ecological trait labels. Second, building upon BioCLIP2, we introduce a two-stage training framework to effectively align audio representations with visual and textual representations. Third, we develop a cross-modal retrieval benchmark that covers all possible directional retrieval across the three modalities (i.e., image-to-audio, audio-to-text, text-to-image, and their reverse directions), with three taxonomic levels: Family, Genus, and Species. Extensive experiments demonstrate that our model learns a unified representation space that captures species-level semantics beyond taxonomy, advancing multimodal biodiversity understanding.
The code is available here.
ZeroPlantSeg: Zero-shot Plant Individual Segmentation
Foundation segmentation models achieve reasonable leaf instance extraction from top-view crop images without training (i.e., zero-shot). However, segmenting entire plant individuals with each consisting of multiple overlapping leaves remains challenging. This problem is referred to as a hierarchical segmentation task, typically requiring annotated training datasets, which are often species-specific and require notable human labor. To address this, we introduce ZeroPlantSeg, a zero-shot segmentation for rosette-shaped plant individuals from top-view images. We integrate a foundation segmentation model, extracting leaf instances, and a vision-language model, reasoning about plants’ structures to extract plant individuals without additional training. Evaluations on datasets with multiple plant species, growth stages, and shooting environments demonstrate that our method surpasses existing zero-shot methods and achieves better cross-domain performance than supervised methods.
The code is available here.

NeuraLeaf: Neural Parametric Leaf Models
We develop a neural parametric model for 3D leaves for plant modeling and reconstruction that are essential for agriculture and computer graphics. While neural parametric models are actively studied for humans and animals, plant leaves present unique challenges due to their diverse shapes and flexible deformation. To this problem, we introduce a neural parametric model for leaves, NeuraLeaf. Capitalizing on the fact that flattened leaf shapes can be approximated as a 2D plane, NeuraLeaf disentangles the leaves’ geometry into their 2D base shapes and 3D deformations. This representation allows learning from rich sources of 2D leaf image datasets for the base shapes, and also has the advantage of simultaneously learning textures aligned with the geometry. To model the 3D deformation, we propose a novel skeleton-free skinning model and create a newly captured 3D leaf dataset called DeformLeaf. We show that NeuraLeaf successfully generates a wide range of leaf shapes with deformation, resulting in accurate model fitting to 3D observations like depth maps and point clouds.
The code is available here.

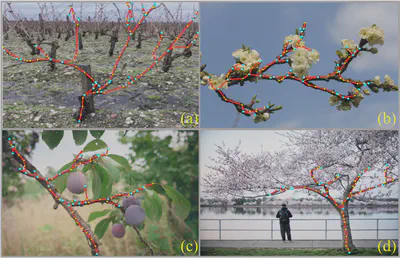
TreeFormer: Plant Skeleton Estimation via Graph Generation
Accurate estimation of plant skeletal structure (e.g., branching structure) from images is essential for smart agriculture and plant science. Unlike human skeletons with fixed topology, plant skeleton estimation presents a unique challenge, i.e., estimating arbitrary tree graphs from images. While recent graph generation methods successfully infer thin structures from images, it is challenging to constrain the output graph strictly to a tree structure. To this problem, we present TreeFormer, a plant skeleton estimator via tree-constrained graph generation. Our approach combines learning-based graph generation with traditional graph algorithms to impose the constraints during the training loop. Specifically, our method projects an unconstrained graph onto a minimum spanning tree (MST) during the training loop and incorporates this prior knowledge into the gradient descent optimization by suppressing unwanted feature values. Experiments show that our method accurately estimates target plant skeletal structures for multiple domains: Synthetic tree patterns, real botanical roots, and grapevine branches.
The code is available here.